Single Protein-Mediated Vesicle Division
Steering group member of AUNAB, Prof. Brigitte Städler, has published a collaborative study in Advanced Materials on the successful reconstitution of the Drs2p–Cdc50p lipid flippase in polymer–lipid hybrid vesicles.
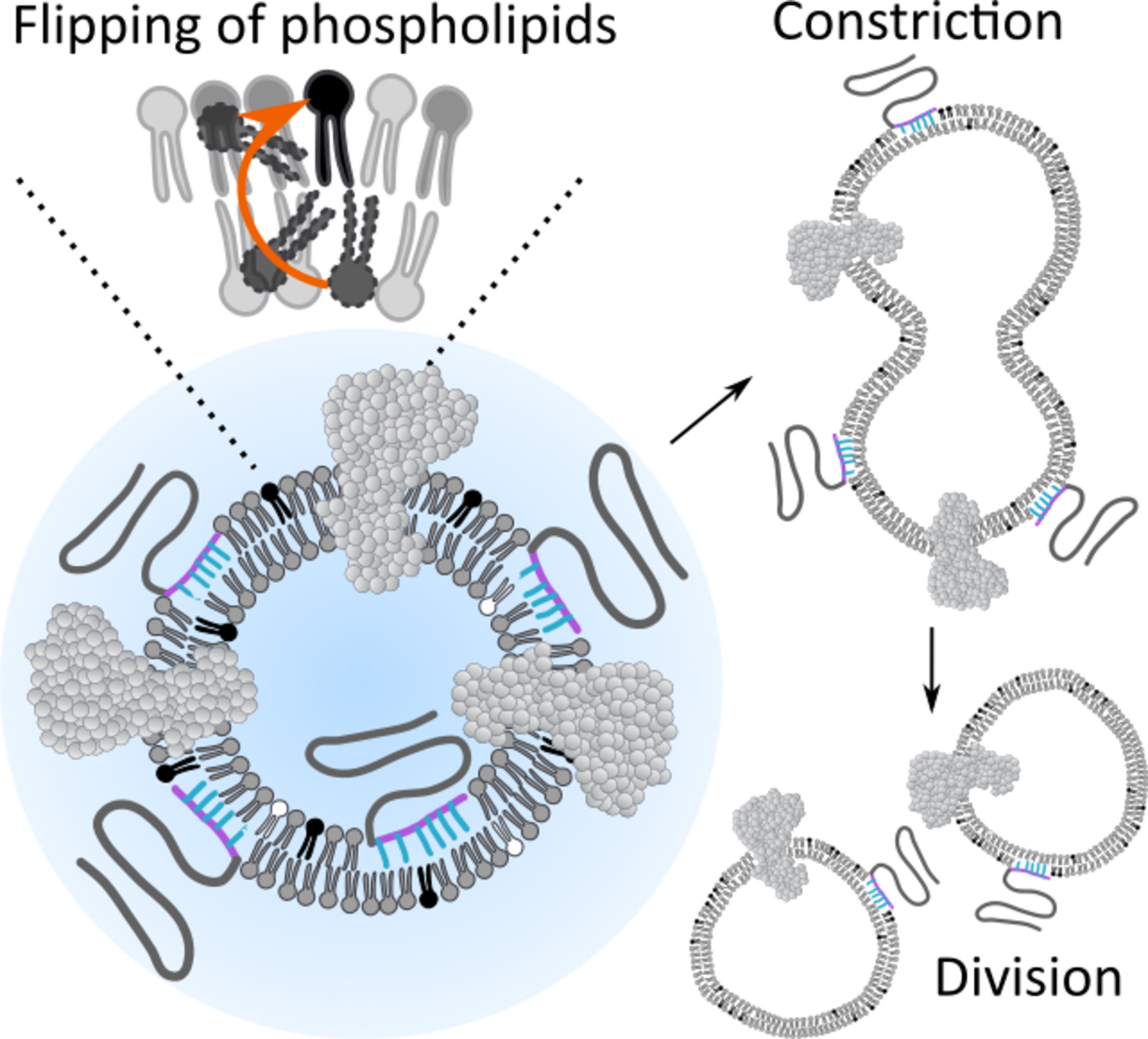
A collaborative study between the Lyons and Stadler labs, published in Advanced Materials, reports the successful reconstitution of the Drs2p–Cdc50p lipid flippase in polymer–lipid hybrid vesicles, enabling active translocation of negatively charged lipids to establish transmembrane asymmetry. Crucially, the chemical nature of the hydrophobic block in the amphiphilic block copolymers determines the curvature changes that drive vesicle constriction and division. This work marks an important step toward constructing synthetic systems capable of protein-mediated division, advancing the goal of bottom-up self-replicating assemblies.
Read the article published in Advanced Marterials here.
Contact information:
Professor Brigitte Maria Städler
Aarhus University
Interdisciplinary Nanoscience Center (iNANO)
Email: bstadler@inano.au.dk
Tenure track Assistant Professor Joseph Lyons
Aarhus University
Interdisciplinary Nanoscience Center (iNANO)
Email: lyons@inano.au.dk
